Cell Hashing is a method that enables sample multiplexing and super-loading on single cell RNA-sequencing platforms, developed in the Technology Innovation lab at the New York Genome Center in collaboration with the Satija lab.
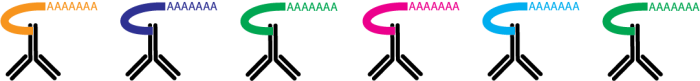
Cell Hashing uses a series of oligo-tagged antibodies against ubiquitously expressed surface proteins with different barcodes to uniquely label cells from distinct samples, which can be subsequently pooled in one scRNA-seq run. By sequencing these tags alongside the cellular transcriptome, we can assign each cell to its sample of origin, and robustly identify doublets originating from multiple samples.
Sample multiplexing schematic:
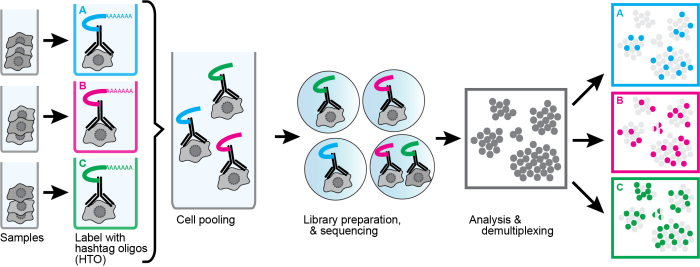
Sample super-loading schematic:
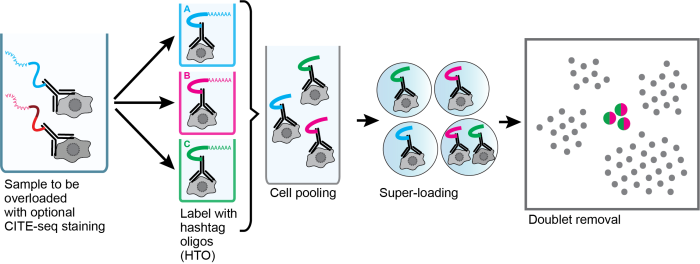
Read more here: Genome Biology paper (December 19, 2018)
Pre-print (December 21, 2017)
Note: The same scheme described for Cell Hashing has been applied to multiplexing nuclei for single nucleus RNA-seq by Jellert Gaublomme, Aviv Regev and collaborators (November 23, 2018). Read about it here