We have adapted CITE-seq and Cell Hashing to be compatible with the 5P / V(D)J single cell kit from 10x Genomics, to allow researchers to perform sample multiplexing, doublet detection and protein detection together with 5’ gene expression and V(D)J reconstruction. In addition, through the addition of a custom RT primer and some additional PCR reactions on the low molecular weight cDNA fraction, we have demonstrated the ability to directly capture sgRNA sequences to perform single cell CRISPR screens with multimodal readout.
Collectively we refer to these capabilities as ECCITE-seq for Expanded CRISPR-compatible CITE-seq. The different modalities can be detected all together, or in desired combinations.
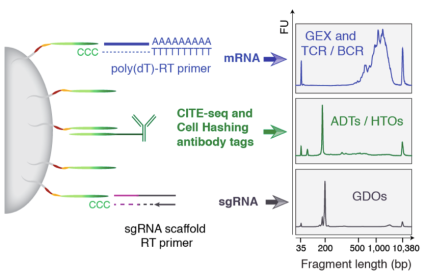
For 5P CITE-seq / 5P Cell Hashing, the oligos conjugated to the antibodies are different than those we have previously described for CITE-seq and Cell Hashing using the 10x 3P kit / Drop-seq / ddSeq.
In place of the polyA stretch on the original CITE-seq / Cell Hashing oligos, 5P CITE-seq and 5P Cell Hashing oligos have a sequence complementary to the 10x Genomics “switch oligo” (see below).
5P CITE-seq antibody-oligos (ADTs):
5P CITE-seq antibody-oligos contain standard TruSeq small RNA read 2 sequences and can be amplified using Illumina’s Truseq Small RNA primer sets (See CITE-seq protocol). See example below with a 12nt barcode (•••) and optional phosphorothioate bonds (*):
5’ CCTTGGCACCCGAGAATTCCA••••••••••••CCCATATAAGA*A*A
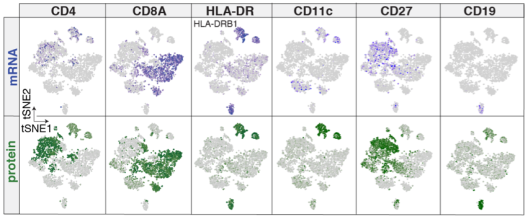
5P Cell Hashing antibody-oligos (HTOs):
5P Cell Hashing antibody-oligos contain standard TruSeq DNA read 2 sequences and can be amplified using truncated versions of Illumina’s TruSeq DNA primer sets (see Cell hashing protocol). See example below with a 12nt barcode (•••) and optional phosphorothioate bonds (*):
5’ GTGACTGGAGTTCAGACGTGTGCTCTTCCGATCT••••••••••••CCCATATAAGA*A*A
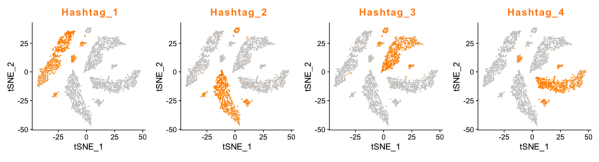
Otherwise, the protocols for both 5P CITE-seq and 5P Cell Hashing remain the same as for 3P CITE-seq and 3P Cell Hashing, performed separately, or together. Antibody-derived tags for both CITE-seq and Cell Hashing are separated from the mRNA-derived cDNA at the same step of the protocol as for the 3P kit, and the same oligos are used for downstream library construction steps.
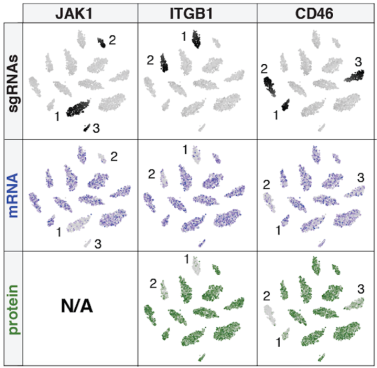
Single guide RNA (sgRNA) capture:
Addition of a custom RT primer that anneals to the sgRNA scaffold enables direct detection of guide sequences, permitting multimodal readout of gene perturbations at the level of single cells.
ASSAY SCHEMES
Please see the assay schemes below for oligo design information, or contact us.
ECCITE_schemes (combines 5P CITE-seq, 5P Cell Hashing and direct guide capture)